Understanding Conformal Prediction
When it comes to understanding Conformal Prediction, everyone has their preferences some like going into the math behind it and research papers, some like looking at code adaptions, and some like reading guides and tutorials like this one.
I’ll approach this article by explaining it in very crude layman’s terms so that the intuition behind Conformal Prediction can be easily grasped and readily applied to your existing and future projects.
Also, I’ll point you toward resources that feature the other approaches mentioned above in the “Where can I learn more about Conformal Prediction” section.
Conformal Prediction usually “works” in the following way:
- Take a trained ML model
- Create an additional set from the data called the calibration set (unseen by the model)
- Pick an error metric (calibration score) and apply a cutoff point (e.g. 95% quantile cutoff point)
- Use the cutoff point to inform the width of your prediction interval
- Use this cutoff point to form the prediction sets for new examples
- Now, most of your predictions should fall inside the prediction interval
That’s it! You now can adapt this for different approaches and models. For example, if you had an image classifier trained to classify dolphins the prediction interval wouldn’t hold labels of more uncertain classes that are being kept away by the chosen quantile level.
Also, the more uncertain the model is (e.g. the harder the inputs are), the larger the prediction set will be and vice-versa which is exactly what we’re looking for. This can be further informed by the average set size, set spread, and coverage that we’ll explore further in this article.
Finally, the error metric (also known as the calibration score) is a very important part and engineering choice that informs everything else when interpreting the results of conformal prediction.
Let’s go into coding so that we can grasp this further.
How to apply Conformal Prediction for Classification with MAPIE?
To apply Conformal Prediction for classification with MAPIE, we will estimate a prediction set of multiple classes such that the probability of a true label of a new test point is always higher than the target confidence level.
We’ll use the classifier’s softmax score output as the conformity score on the Iris dataset. The classifier will be the Naive Bayes classifier from the scikit-learn library.
To execute this properly, we’ll need to follow these steps:
- Craft a toy dataset and form train, calibration, and test sets.
- Fit the model on the train set.
- Set the conformal score to be the softmax output of the true class for each sample in the calibration set
- Define
as being the quantile of calibration scores (S1,…,Sn) which is
- Form a prediction set for each new test data point (where Xn+1 is known but Yn+1 isn’t) so that it includes all the labels with a sufficiently high softmax output.
Prior to doing the first two steps, we will import the libraries we need:
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from sklearn.model_selection import train_test_split
from sklearn import naive_bayes
from mapie import MapieClassifier
from mapie.metrics import classification_coverage_scoreNow, the first two steps:
# Create a toy dataset with 2 features and 3 classes (0, 1, 2) with a bit of noise
n_samples = 1500
n_features = 2
n_classes = 3
X = np.random.randn(n_samples, n_features)
y = np.zeros(n_samples)
for i in range(n_classes):
X[y == i] += np.random.randn(1, n_features) * 1.2
y = np.where(X[:, 0] > 0, 0, 1)
y = np.where(X[:, 1] > 0, y, y + 1)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1, random_state=42)
X_train, X_cal, y_train, y_cal = train_test_split(X_train, y_train, test_size=0.1, random_state=42)
# Train a Gaussian Naive Bayes classifier
clf = naive_bayes.GaussianNB()
clf.fit(X_train, y_train)Before moving on, let’s plot our training set data:
plt.figure(figsize=(7, 7))
plt.scatter(X_train[:, 1], X_train[:, 2], c=y_train, s=100, cmap='viridis', edgecolor='k', linewidth=2)
plt.xlabel('Feature 1')
plt.ylabel('Feature 2')
plt.show()Okay, now we proceed with computing the probabilities on the test set, computing the softmax scores, and passing the classifier as the estimator for MapieClassifier. Then we can fit the MapieClassifier on the calibration sets.
Here also comes the part where we pick the alpha metric (alpha = 1 - target coverage) which will suggest the quantile we are to set. In other words, the lower the alpha is, the higher the quantile will be. The higher the quantile, the less likely that a score satisfies it in highly uncertain areas.
Let’s pick 3 different numbers for alpha so that we can visualize the difference it makes.
# Predict class labels and probabilities
y_pred_proba = clf.predict_proba(X_test)
# Calculate Softmax score
softmax_score = np.max(y_pred_proba, axis=1)
# Initialize the Conformal Prediction classifier
mapie_score = MapieClassifier(estimator=clf, cv="prefit", method="score")
mapie_score.fit(X_cal, y_cal)
alpha = [0.2, 0.1, 0.05]
y_pred_score, y_ps_score = mapie_score.predict(X_test, alpha=alpha)method="score"– is based on the scores (i.e. 1 minus the softmax score of the true label) on the calibration sety_pred_score– represents the prediction in the test set by the base estimator.y_ps_score– represents the prediction sets estimated by MAPIE.
To visualize the differences between different alpha scores, I’ll use a function kindly provided inside the MAPIE documentation:
def plot_scores(n, alphas, scores, quantiles):
colors = {0: "#1f77b4", 1: "#ff7f0e", 2: "#2ca02c"}
plt.figure(figsize=(7, 5))
plt.hist(scores, bins="auto")
for i, quantile in enumerate(quantiles):
plt.vlines(
x=quantile,
ymin=0,
ymax=400,
color=colors[i],
ls="dashed",
label=f"alpha = {alphas[i]}"
)
plt.title("Distribution of scores")
plt.legend()
plt.xlabel("Scores")
plt.ylabel("Count")
plt.show()Prior to using the function, we need to obtain the scores and quantiles from MAPIE. Then, we can plot the difference:
scores = mapie_score.conformity_scores_
n = len(mapie_score.conformity_scores_)
quantiles = mapie_score.quantiles_
plot_scores(n, alpha, scores, quantiles)In the graph above we can observe how different alpha values affect the quantile and the overall width of our conformal prediction sets. We can also observe that the spread looks good and we’ll double down on this by exploring the coverage of each alpha score.
Before that, it is worth repeating that a high alpha might lead to a high quantile that might not be reached by any class in regions where the uncertainty is high. For example, this would be the region where there is an overlap between the classes. Especially the very middle of the chart.
Now, let’s plot the differences for different values of alpha. I’ll use the plotting function that is provided inside the MAPIE docs and adapt it to not show a mesh but rather individual points.
On the top left, you will see the prediction made by the base estimator. The other 3 subplots will feature each alpha score.
Points that are red had high uncertainty and the model abstained from giving a prediction for them (lower coverage), while points that are highly satiated in purple shared multiple possible classes in the prediction set (higher coverage).
plot_results(alpha, X_test, y_pred_score, y_ps_score)Take note that there are alterations to this method (e.g. Adaptive Prediction Set) whose conformity score is the cumulated score of the softmax output until the true label is reached. This is for those that don’t want null predictions under high uncertainty.
You have probably raised an important issue here. How does one pick the right alpha score? Well, to do this, MAPIE provided us with an example of testing a wide range of alpha values and observing their coverage scores and average prediction set sizes.
The full code can be found in the GitHub repository and here’s the output featuring the graph of different alpha score coverages and average set sizes for each alpha score:
How to apply Conformal Prediction for Regression with MAPIE?
To apply Conformal Prediction for Regression with MAPIE, we will use a conformalized quantile regression estimator (LGBMRegressor) with the MapieQuantileRegressor module on example tabular data.
Have in mind that you can use the MapieRegressor for other regression types that aren’t quantile regressors. Also, some code will be hidden for brevity and can be found inside the GitHub repository.
The steps that we’ll do are the following:
- Load the Diamonds dataset and preprocess it (we remove the categorical variables for simplicity)
- Split the dataset into train, calibration, and test sets
- Optimize and train the Quantile LGMBRegressor
- Pass the estimator to MAPIE and fit it with 80% coverage (alpha=0.2)
- Plot the conformal prediction sets and errors
Let’s perform the first two steps:
# import the diamonds dataset
df = pd.read_csv("https://raw.githubusercontent.com/mwaskom/seaborn-data/master/diamonds.csv")
# Create a train, calibration and test set
X = df.drop(["price"], axis=1)
# drop categorical variables for brevity
X = X.drop(["cut", "color", "clarity"], axis=1)
y = df["price"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
X_cal, X_test, y_cal, y_test = train_test_split(X_test, y_test, test_size=0.5, random_state=42)To observe the distribution of diamond prices we can do the following:
plt.hist(y_train, bins="auto")
plt.title("Distribution of prices")
plt.xlabel("Price")
plt.ylabel("Count")
plt.show()Now, we optimize the model:
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import KFold
# fit a Quantile LightGBM model
clf = LGBMRegressor(
objective="quantile",
alpha=0.05,
random_state=42
)
# optimize the model
params_distributions = dict(
num_leaves=np.random.randint(10, 50, 5),
max_depth=np.random.randint(3, 20, 1),
n_estimators=np.random.randint(50, 300, 10),
learning_rate=np.random.uniform(0.01, 0.1, 10),
)
optim_model = RandomizedSearchCV(
clf,
param_distributions=params_distributions,
n_jobs=-1,
n_iter=100,
cv=KFold(n_splits=5, shuffle=True),
verbose=-1
)
optim_model.fit(X_train, y_train)
clf = optim_model.best_estimator_After this, we pass the model onto MAPIE:
mapie = MapieQuantileRegressor(
clf,
method="quantile",
cv="split",
alpha=0.2
)
mapie.fit(X_train, y_train)
# sort the data
y_pred, y_pis = mapie.predict(X_test)
(
y_test_sorted,
y_pred_sorted,
lower_bound,
upper_bound
) = sort_y_values(y_test, y_pred, y_pis)
# Calculate the coverage and width of the prediction intervals
coverage = regression_coverage_score(
y_test,
y_pis[:, 0, 0],
y_pis[:, 1, 0]
)
width = regression_mean_width_score(
y_pis[:, 0, 0],
y_pis[:, 1, 0]
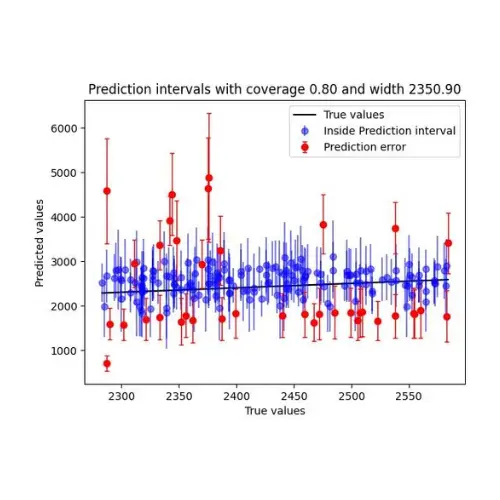
)Now, we can plot our prediction intervals, the true value, and the error intervals. The prediction interval will be plotted where the true value was inside of it, and the error interval will be the interval inside of which the true value wasn’t present.
I’ll only plot these for a small subset of the test set as the graph can get messy. I encourage everyone to experiment with different parts of the test set to observe the difference. For example, the model performs better on medium-to-high prices than on lower ones.
As the prices go up, so do our prediction intervals.
How to perform forecasting with Conformal Prediction in Python?
To perform forecasting with Conformal Prediction in Python, you can use any forecasting algorithm and create training and test sets. Then, use a library such as MAPIE to calibrate the algorithm.
You can also explore the option of coding your own conformal prediction implementations.
Below, I’ll show an example of a monthly sales dataset I’ve found online. It will come with the GitHub repo.
After creating the dataset we will proceed by engineering a few features and splitting it into the train and test sets. We also load in the MapieRegressor and XGBoost as our algorithms of choice.
from mapie.time_series_regression import MapieTimeSeriesRegressor
from mapie.subsample import BlockBootstrap
import xgboost as xgb
df = pd.read_csv("sales.csv")
df["date"] = pd.to_datetime(df["date"])
df = df.set_index("date")
# create features
df["month"] = df.index.month
df["year"] = df.index.year
# plot the data
plt.plot(df["sales"])
plt.title("Sales")
plt.xlabel("Date")
plt.ylabel("Sales")Now, let’s make this more fun by introducing a sudden change to the time series to see how the model will react. We’ll first split it into 80% train and 20% test after which we introduce an abrupt change in the label at the start of 1975 by applying a 10% drop rate.
# create train and test sets
X = df.drop(["sales"], axis=1)
y = df["sales"]
# split the data where test is eveyrthing after 1974
X_train = X[X.index.year < 1974]
X_test = X[X.index.year >= 1974]
y_train = y[y.index.year < 1974]
y_test = y[y.index.year >= 1974]
# introduce a change in seasonality after 1975 in the y_test set by applying a 10% drop
y_test.loc[y_test.index.year >= 1975] = y_test.loc[y_test.index.year >= 1975] * 0.9
# plot the data whole data and colored train and test sets
plt.plot(X_train.index, y_train, label="Train set")
plt.plot(X_test.index, y_test, label="Test set")
plt.title("Sales")
plt.xlabel("Date")
plt.ylabel("Sales")
plt.legend()Due to the introduction of the abrupt change, I expect the algorithm to struggle to capture the change. Here, we hope that conformal prediction will be able to adjust for the change by having the true value inside its prediction sets.
Now, we will train the XGBoost model and optimize its hyperparameters:
reg = xgb.XGBRegressor(
n_estimators=1000,
random_state=42
)
# optimize the model
params_distributions = dict(
max_depth=np.random.randint(3, 20, 1),
learning_rate=np.random.uniform(0.01, 0.1, 10),
)
reg = RandomizedSearchCV(
reg,
param_distributions=params_distributions,
n_jobs=-1,
n_iter=10,
cv=KFold(n_splits=5, shuffle=True),
verbose=-1
)
reg.fit(X_train, y_train)Now, we can estimate the prediction intervals by using the MapieTimeSeriesRegresor. Here, we have two options to estimate the prediction intervals:
- the regular .fit and .predict process, limiting the use of training set residuals to create prediction intervals
- the use of
.partial_fitin addition to.fitand.predictallowing MAPIE to use new residuals from the test points as new data are becoming available.
The second approach is better for time series with unexpected oscillations. I’ll show you both versions so that you can see the difference between the two.
Take note that we’ll be using the BlockBootstrap method which is shown to be better for time-series modeling. As for our coverage, we will go with 95% by having the alpha be 0.05. The method for the MAPIE regressor will be EnbPi which was shown to perform well on time series data.
# It's Confromal Prediction time!
alpha = 0.05
gap = 1
mapie_cv = BlockBootstrap(
n_blocks=10,
overlapping=True,
random_state=42
)
mapie_enbpi = MapieTimeSeriesRegressor(
reg,
method="enbpi",
cv=mapie_cv,
agg_function="mean",
n_jobs=-1
)Now, we start by applying the first method:
mapie_enbpi = mapie_enbpi.fit(X_train, y_train)
y_pred_npfit, y_pis_npfit = mapie_enbpi.predict(
X_test, alpha=alpha, ensemble=True, optimize_beta=True
)
coverage_npfit = regression_coverage_score(
y_test, y_pis_npfit[:, 0, 0], y_pis_npfit[:, 1, 0]
)
width_npfit = regression_mean_width_score(y_pis_npfit[:, 0, 0], y_pis_npfit[:, 1, 0])Now, we apply the second method:
mapie_enbpi = mapie_enbpi.fit(X_train, y_train)
y_pred = np.zeros((len(test), 2))
y_pis = np.zeros((len(test), 2, 2))
conformity_scores_pfit = []
lower_quantiles_pfit = []
higher_quantiles_pfit = []
y_pred[:gap], y_pis[:gap, :, :] = mapie_enbpi.predict(
X_test.iloc[:gap, :], alpha=alpha, ensemble=True, optimize_beta=True
)
for step in range(gap, len(X_test), gap):
mapie_enbpi.partial_fit(
X_test.iloc[(step - gap) : step, :],
y_test.iloc[(step - gap) : step],
)
(y_pred[step : step + gap], y_pis[step : step + gap, :, :],) = mapie_enbpi.predict(
X_test.iloc[step : (step + gap), :],
alpha=alpha,
ensemble=True,
optimize_beta=True,
)
conformity_scores_pfit.append(mapie_enbpi.conformity_scores_)
lower_quantiles_pfit.append(mapie_enbpi.lower_quantiles_)
higher_quantiles_pfit.append(mapie_enbpi.higher_quantiles_)
coverage_pfit = regression_coverage_score(y_test, y_pis[:, 0, 0], y_pis[:, 1, 0])
width_pfit = regression_mean_width_score(y_pis[:, 0, 0], y_pis[:, 1, 0])Finally, when we plot the prediction intervals and get the following chart:
The coverage in both examples isn’t great, but we can see how the introduction of the partial fit managed to “recover” after the abrupt change in the label by adjusting its prediction intervals.
Please feel free to explore different alpha values and how they affect the output intervals of the algorithm.